ePolyScat advanced usage tutorial
08/05/20
Paul Hockett
Disclaimer: I am an enthusiastic ePolyScat user for photoionization calculations, but not an expert on the code. Nonetheless, this tutorial aims to go over some of the key features/uses of ePS for such problems - as far as my own usage goes - and provide an introduction and resource to new users.
Overview & resources
ePolyScat (ePS) is an open-source tool for numerical computation of electron-molecule scattering & photoionization by Lucchese & coworkers. For more details try:
The ePolyScat website and manual. Note that the manual uses a frames-based layout, use the menu on the page to navigate to sub-sections (direct links break the menu).
Calculation of low-energy elastic cross sections for electron-CF4 scattering, F. A. Gianturco, R. R. Lucchese, and N. Sanna, J. Chem. Phys. 100, 6464 (1994), http://dx.doi.org/10.1063/1.467237
Cross section and asymmetry parameter calculation for sulfur 1s photoionization of SF6, A. P. P. Natalense and R. R. Lucchese, J. Chem. Phys. 111, 5344 (1999), http://dx.doi.org/10.1063/1.479794
Applications of the Schwinger variational principle to electron-molecule collisions and molecular photoionization. Lucchese, R. R., Takatsuka, K., & McKoy, V. (1986). Physics Reports, 131(3), 147–221. https://doi.org/10.1016/0370-1573(86)90147-X (comprehensive discussion of the theory and methods underlying the code).
ePSproc is an open-source tool for post-processing & visualisation of ePS results, aimed primarily at photoionization studies.
Ongoing documentation is on Read the Docs.
Source code is available on Github.
For more background, see the software metapaper for the original release of ePSproc (Aug. 2016): ePSproc: Post-processing suite for ePolyScat electron-molecule scattering calculations, on Authorea or arXiv 1611.04043.
ePSdata is an open-data/open-science collection of ePS + ePSproc results.
ePSdata collects ePS datasets, post-processed via ePSproc (Python) in Jupyter notebooks, for a full open-data/open-science transparent pipeline.
ePSdata is currently (Jan 2020) collecting existing calculations from 2010 - 2019, from the femtolabs at NRC, with one notebook per ePS job.
In future, ePSdata pages will be automatically generated from ePS jobs (via the ePSman toolset, currently in development), for immediate dissemination to the research community.
Source notebooks are available on the Github project pages, and notebooks + datasets via Zenodo repositories (one per dataset). Each notebook + dataset is given a Zenodo DOI for full traceability, and notebooks are versioned on Github.
Note: ePSdata may also be linked or mirrored on the existing ePolyScat Collected Results OSF project, but will effectively supercede those pages.
All results are released under Creative Commons Attribution-NonCommercial-ShareAlike 4.0 (CC BY-NC-SA 4.0) license, and are part of our ongoing Open Science initiative.
Workflow
The general workflow for photoionization calculations plus post-processing is shown below. This pipeline involves a range of code suites, as shown in the main workflow; some additional details are also illustrated.
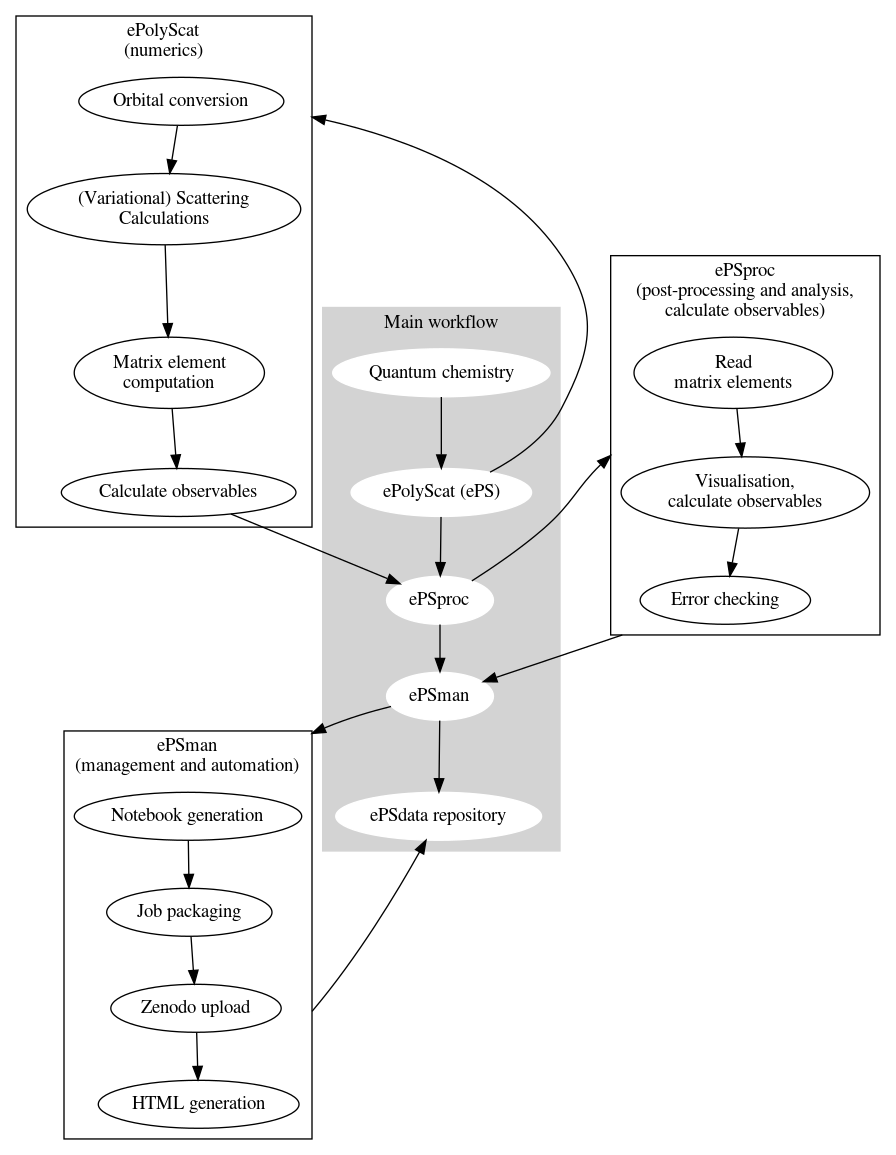
ePS advanced usage
As indicated in the basic usage notebook, there are a range of commands in ePS for additional computations and outputs. Explore the Commands page of the manual for more details.
In this tutorial these capabilities won’t be explored further, aside from the to note two particularly interesting commands:
DumpIdy: write photoionization matrix elements to output fileViewOrb: write (gridded) wavefunction to file
In particular, the DumpIdy command provides a route to further post-processing/interfacing to other codes.
ePSproc has limited support for ViewOrb, with further support planned for the near future. Here’s an example output, for the ABCO molecule (with legacy ePSproc Matlab code):
ePSproc basic usage
ePSproc implements some basic parsing of ePS files, plotting and further post-processing. Currently:
CrossSectionsegments can be read and plotted.DumpIdymatrix element segments can be read and plotted.Further properties (AF-PADs, MF-PADs etc) can be computed from the matrix elements.
Example notebooks can be found in the ePSproc docs:
(TODO - add some demo material directly here.)
Working with dipole matrix elements
Why work with matrix elements?
Working directly from the dipole matrix elements has a few benefits, mainly relating to the flexibility these provide for post-processing and further use:
Fundamental molecular properties, these can be compared with experimentally obtained values (see platfrom development for quantum metrology with photoelectrons notes).
Analysis of partial wave composition (also available directly from ePS using
GetCrowith the CroByPartialWave option set).Provides a minimal dataset which can be piped into further analysis/processing routines.
Computation of additional properties based on the matrix elements, e.g. molecular frame (MF) and aligned frame (AF) observables. (Some of these are already possible in ePS directly, e.g.
OrientCrofor MF computations).Open-science: dissemination and sharing of results & processing (see ePSdata).
Disadvantages…
Care required for consistency and correct treatment (e.g. phase conventions, normalisation, etc.).
Additional formalism and derivations required.
ePSproc: workflow with matrix elements
The basic ePSproc workflow (from the original software metapaper) is shown below: essentially, the ePS output file is parsed for photoionization matrix elements, and ancillary data; the matrix elements are the used to calculate properties of interest, such as photoionziation cross-sections and MF-PADs. For ePSdata, this workflow is applied to each ePS dataset, with a Jupyter notebook as a template. The completed analysis & dataset is then uploaded to Zenodo, and HTML version to ePSdata.
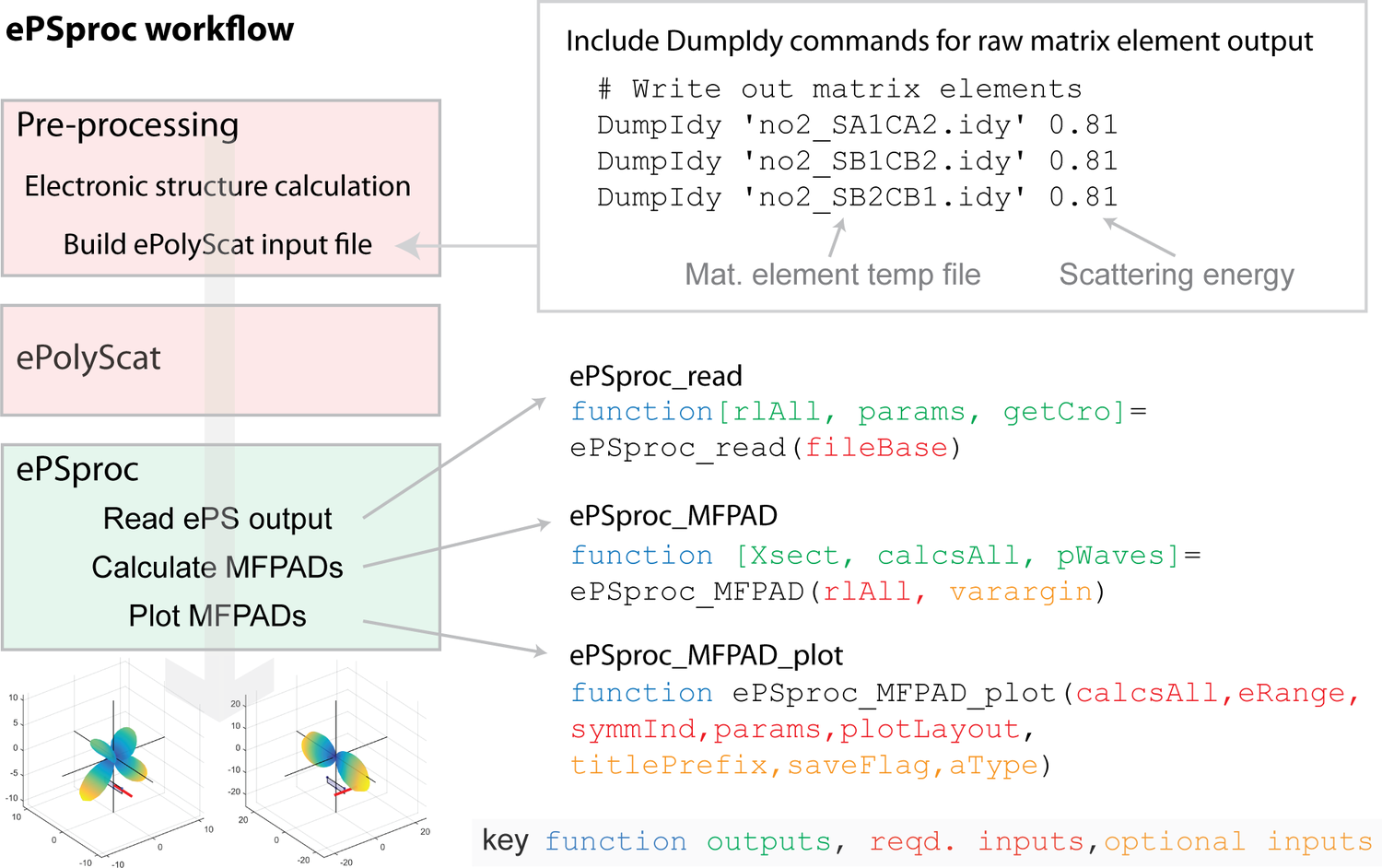

For further usage examples, see:
… and the results on ePSdata.